Visualising protein interaction
Professor Juliet Gerrard, Michael Barnett, PhD Candidate, School of Biological Sciences; David Doak, Norwich University of Arts, UK; Nick Young, Bianca Haux , Centre for eResearchBackground
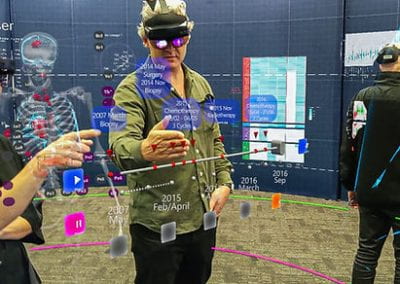
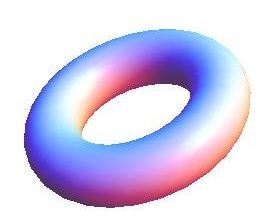
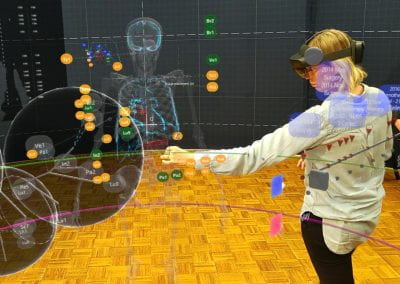
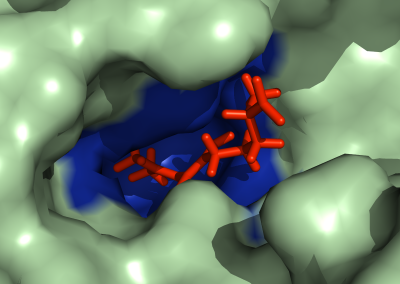
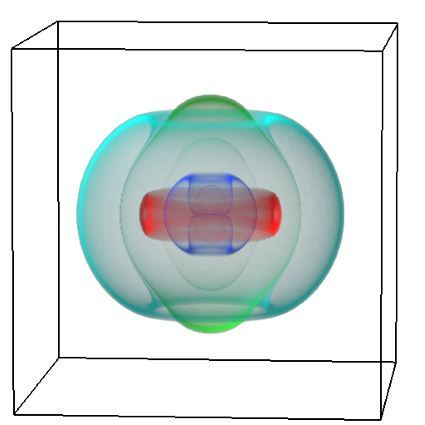
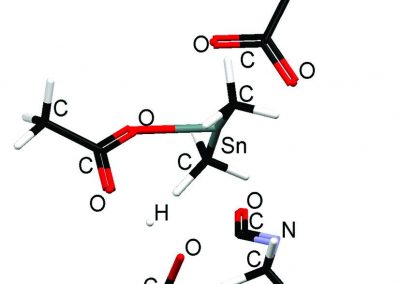
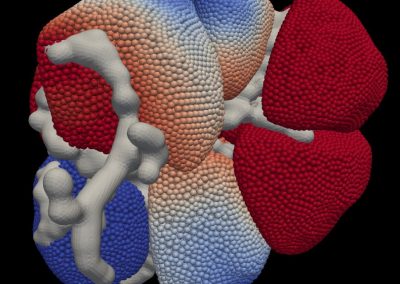
Auckland molecular research at the School of Biological Sciences aims to build capability in the use of virtual reality technology for molecular visualisation, i.e. to create an experience which enables the user to actually climb inside a molecule and walk around it. For this particular project, the Centre for eResearch (CeR) has been working with Professor Juliet Gerrard and PhD Candidate Michael Barnett from the School of Biological Science’s to visualise the protein bonding of Human Peroxiredoxin 3, the second most common protein in the human body behind Haemoglobin.
Simulation
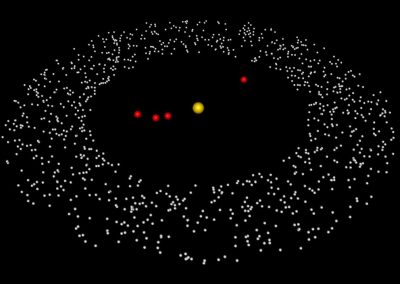
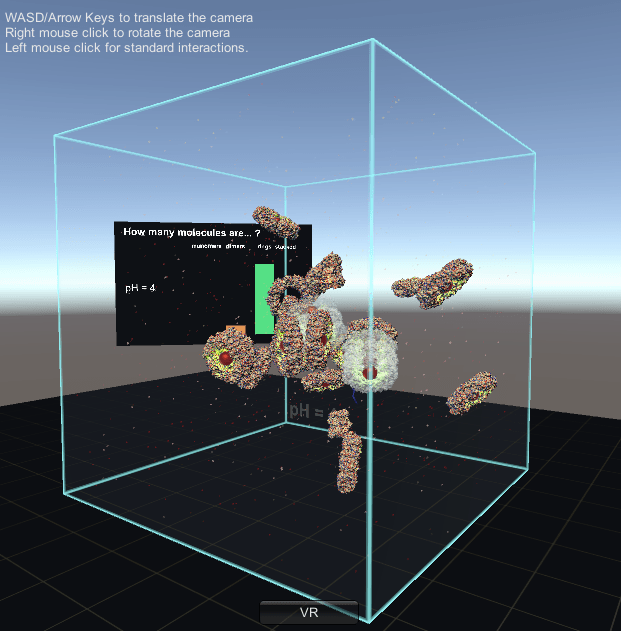
In a solution of PRX3 monomers (single units), the monomers will bond together to form dimers (paired units). These dimers then bond together to form rings, and the rings bond together to form a filament. This entire process has been simulated using the game engine Unity and works in te Vive by Nick Young and Bianca Haux from CeR, in collaboration with Dr David Doak from Norwich University of the Arts, UK.
Self assembly of peroxiredoxi
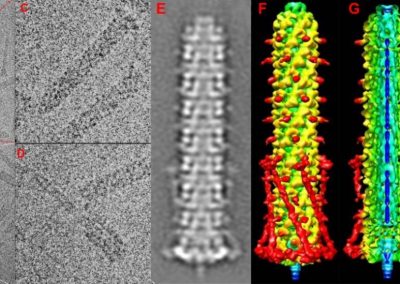
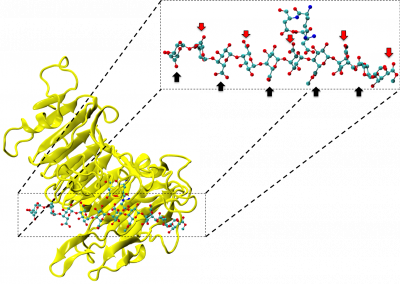
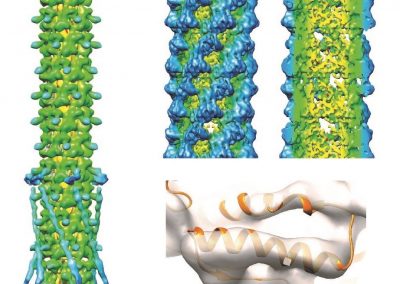
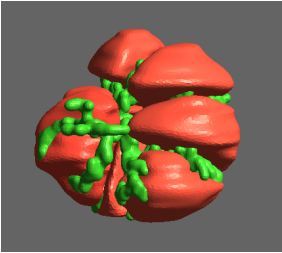
Peroxiredoxins (Prxs, EC 1.11.1.15; pronounced per-oxy- red-ox-in) are a ubiquitous family of highly abundant antioxidant enzymes that protect all cells from free radical damage. They have an intriguing repertoire of self-assembled structures which helps modulate their activity inside cells and also provides an exciting scaffolding opportunity in vitro for nanotechnology applications.
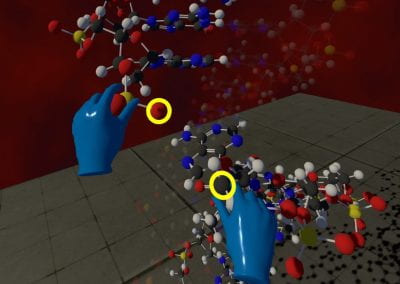
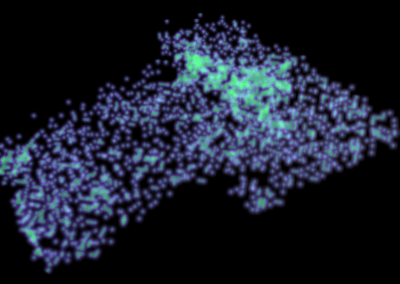
This visualisation shows the assembly of monomers into dimers, dimers into rings, and rings into nanotubes. the user can teleport with either touchpad, and you can adjust the pH by grabbing the handle above one of your controllers with the opposite controller (using the trigger on the underside) and moving it left or right. The pH controls the degree of assembly. You can pick up a monomer, dimer or ring, and can break it into its smaller components by pulling it apart with two hands or shaking it while holding it (if it’s a dimer or a ring). The rings can be re-ordered, or re-pair monomers by simply moving them closer – as monomers/dimers/rings will aim to pair with the closest object of the same type. The project is an open source at https://github.com/uoa-eresearch/prx_fishtank
Further refinement on user experience
The CeR specialists have further done some refinements in order to improve the user experience by adding the following features:
Haptic/tactile feedback to indicate attraction forces feedback.
Added special effects to demonstrate more intermolecular and intramolecular events such as the accretion of free radicals within the peroxiredoxin rings.
Added user interface improvements to allow the users to navigate the program easier.
Added keyboard controls for demo controller for the purpose of allowing for smoother demonstration experience.
Tractor beam function to allow for extra ease of interaction with the molecules to improve the stationary experience while using the app.
Added sound effects to enhance the immersion and emphasis on the events within the application/simulation.
Improvements to the quality of life of the application such as small value tweaks and graphics tweaks to visually improve the app.
Other miscellaneous
There has been a “party mode” added which can be played as a competitive game while at the same time being educating.
See more case study projects:

Our Voices: using innovative techniques to collect, analyse and amplify the lived experiences of young people in Aotearoa
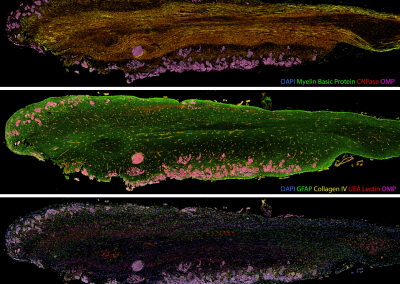
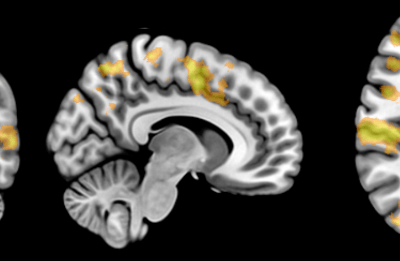
Painting the brain: multiplexed tissue labelling of human brain tissue to facilitate discoveries in neuroanatomy

Detecting anomalous matches in professional sports: a novel approach using advanced anomaly detection techniques
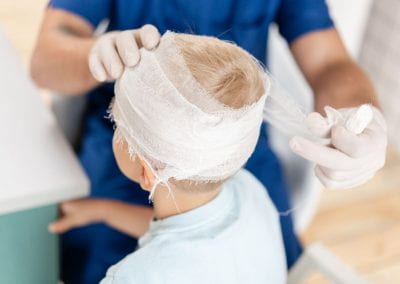
Benefits of linking routine medical records to the GUiNZ longitudinal birth cohort: Childhood injury predictors
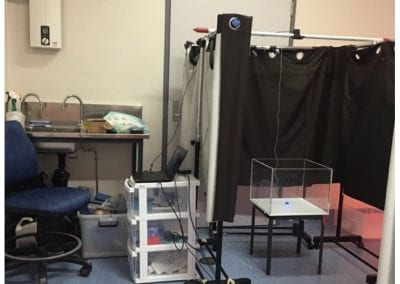
Using a virtual machine-based machine learning algorithm to obtain comprehensive behavioural information in an in vivo Alzheimer’s disease model
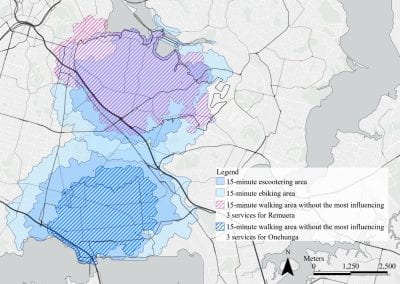
Mapping livability: the “15-minute city” concept for car-dependent districts in Auckland, New Zealand
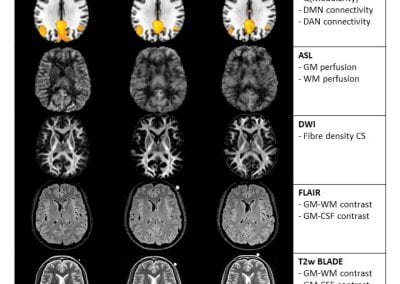
Travelling Heads – Measuring Reproducibility and Repeatability of Magnetic Resonance Imaging in Dementia
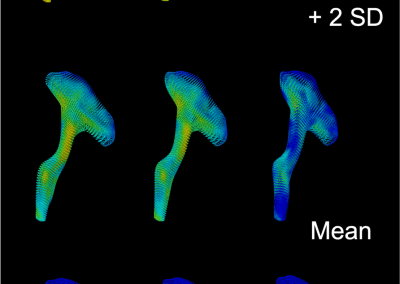
Novel Subject-Specific Method of Visualising Group Differences from Multiple DTI Metrics without Averaging

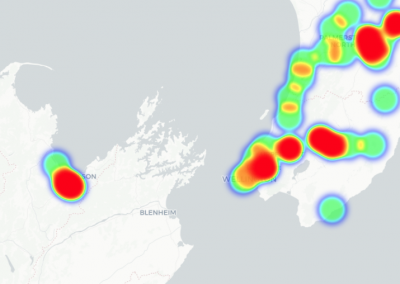
Re-assess urban spaces under COVID-19 impact: sensing Auckland social ‘hotspots’ with mobile location data
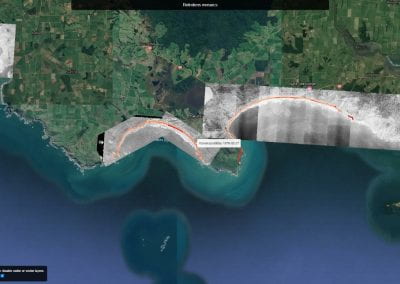
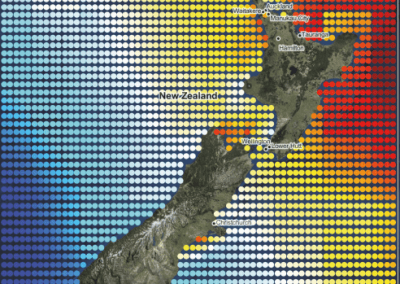
Aotearoa New Zealand’s changing coastline – Resilience to Nature’s Challenges (National Science Challenge)
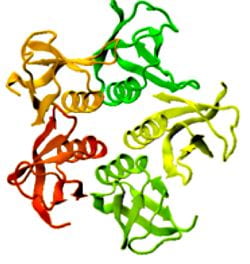
Proteins under a computational microscope: designing in-silico strategies to understand and develop molecular functionalities in Life Sciences and Engineering
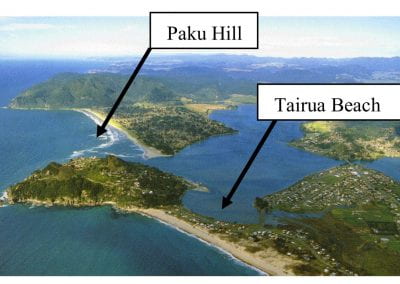
Coastal image classification and nalysis based on convolutional neural betworks and pattern recognition
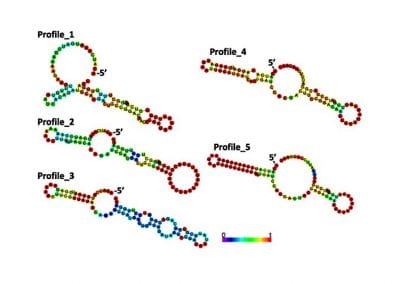
Determinants of translation efficiency in the evolutionarily-divergent protist Trichomonas vaginalis
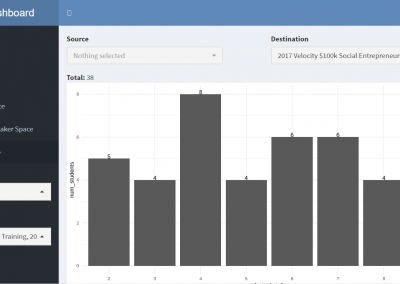
Measuring impact of entrepreneurship activities on students’ mindset, capabilities and entrepreneurial intentions

Using Zebra Finch data and deep learning classification to identify individual bird calls from audio recordings
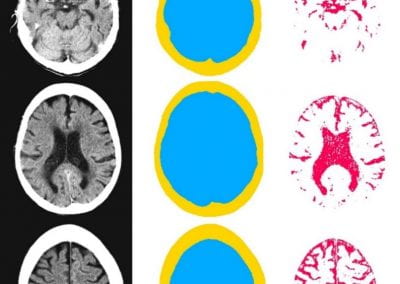
Automated measurement of intracranial cerebrospinal fluid volume and outcome after endovascular thrombectomy for ischemic stroke

Using simple models to explore complex dynamics: A case study of macomona liliana (wedge-shell) and nutrient variations
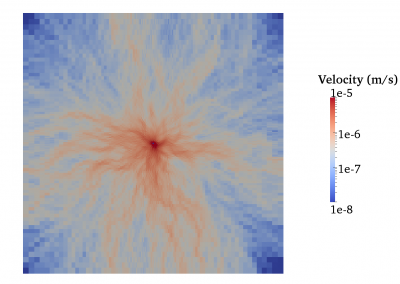
Fully coupled thermo-hydro-mechanical modelling of permeability enhancement by the finite element method

Modelling dual reflux pressure swing adsorption (DR-PSA) units for gas separation in natural gas processing
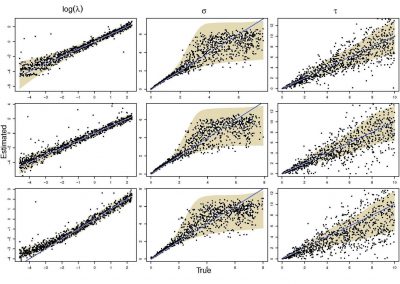
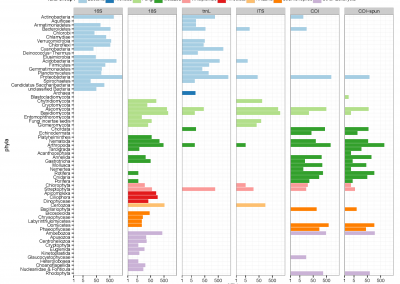
Molecular phylogenetics uses genetic data to reconstruct the evolutionary history of individuals, populations or species
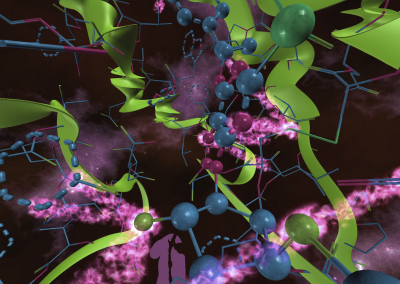
Wandering around the molecular landscape: embracing virtual reality as a research showcasing outreach and teaching tool